Beware the Switch Flippers, Part III
A comprehensive analysis of the interactions of all known human microRNAs with Pfizer, Moderna, SARS-CoV-2 and SARS-CoV spike protein sequence
“Creativity is not accomplished by the intellect, but by the play of instinct” — Carl Jung
One of the great things about mRNA gene therapy (if you happen to be so inclined to develop such products) is that it gives you a certain degree of sophisticated control over the expression of both the “therapeutic” protein and other proteins in the host cell. That control is quite important when the delivery vehicle is a lipid nanoparticle (LNP), which can enter any cell it happens to encounter. Moreover, when the payload is a pathogenic protein, like the SARS-CoV-2 spike, you really need to have some way for the mRNA you are delivering to “sense” where it is in the body to determine if expression is appropriate or if it might be catastrophic. For instance, you really don’t want uncontrolled spike protein expression in the heart and brain.
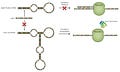
So, how do you go about designing a smart “vaccine”? It’s very simple. All you need to do is design your mRNA sequence to contain one or more target sites that will bind to microRNAs that are upregulated in the body in response to the stressful conditions induced by the presence of a pathogenic substance like the spike protein. The image below shows the basic idea. The microRNA labelled miR-x represents a microRNA that is upregulated during say, myocarditis, which is able to bind to the spike protein mRNA at one of its target sites and subsequently prevent translation of the spike protein by the ribosome.
The process depicted above is just a very simple logical operation that can be represented by a single NOR gate:
So, if any of the miRs shown above happen to be present in the host cell and bind with its corresponding site on the spike protein mRNA, translation will be shut-off. This is smart. You want your “vaccine” to shut-off if it starts pushing the victim too close to acute, catastrophic outcomes.
But wait, we can make the “vaccine” even smarter. What if you’d also like to exert some control over gene expression in the host cell for god knows whatever reason(s)? Well, if you want to target certain genes then just load your mRNA sequence with tons of binding sites for the microRNAs that target that gene’s mRNA. A broad-spectrum approach might be appropriate here, as most genes can be targeted by multiple microRNAs which may be present at differing concentrations in different host cells. You want to soak up as many different microRNAs as you can to upregulate expression of the target gene.

By acting as a sponge for specific microRNAs, the spike protein mRNA can shut-off its own expression in favor of expression of the host cell mRNA that is also targeted by the same microRNAs. Our logic circuit now looks something like this:
Now we’re cooking! By applying just a bit of thought to our “vaccine” design, we’ve endowed it with the ability to perform crude calculations to determine protein expression pathways and also shutdown translation of the spike protein in the event of an emergency.
To summarize, the process from cell entry of the LNP to protein translation is shown below. Once the spike protein mRNA spills out of the LNP into the cell cytoplasm, protein translation can follow one of the three indicated pathways. Paths 1 and 2 show the competition between the spike mRNA and host cell mRNA for a particular microRNA (miR-y). In the first path, the microRNA binds to the spike mRNA and inhibits its translation, while the host cell mRNA can proceed. This path becomes more favorable as the number of binding sites on the spike mRNA increases and/or the number of spike mRNAs in the cell increases. The second path shows the binding of the microRNA to the host cell mRNA which acts to inhibit its translation, while spike mRNA translation can proceed (so long as it hasn’t been bound to a different microRNA). Finally, Path 3 represents the direct binding of a microRNA (miR-x) to the spike protein mRNA which prevents its translation by the ribosome. This case might represent an emergency shut-off caused by the accumulation of specific microRNAs in response to stressful conditions.
My initial efforts on understanding the targeting of microRNAs by the Pfizer and Moderna vaccines were very narrowly focused on a few specific microRNAs. I recently realized that a better approach to this problem might be to just cast a wide net and write a script to analyze the interaction of every known human microRNA with the vaccine sequences. So, that’s what I did. I believe the results are quite compelling.
Method
The workflow I used to perform this analysis is shown in the figure below. First, I collected the spike protein sequences for the Pfizer BNT-162b2 and Moderna mRNA-1273 vaccines from Github. For reference, I also downloaded the sequences for the spike proteins of SARS-CoV-2 Wild-type Wuhan strain (NC_045512.2), SARS-CoV Tor2 strain (NC_004718.3) and HCoV-229E (NC_002645.1). The next step was to collect the sequences for all of the known human microRNAs from miRBase.
The target sequences and microRNA sequences were then fed into the RNA22v2 algorithm to identify high-confidence binding sites. The algorithm was set at the lowest sensitivity (21%) and highest specificity (92%) to exclude low confidence hits. The output from the target calculations was filtered so that only microRNA target sites with a p-value less than or equal to 0.05 were used for further analysis.
The microRNAs with binding sites on the target sequences were then fed into the miRDB for gene target identification. Only the top 10 genes targeted by each microRNA were considered for further analysis.
Finally, the target genes identified in the previous step were fed into the Protein Atlas to extract mRNA expression levels by tissue.
Results
Pfizer and Moderna sequences contain significantly more microRNA binding sites than the reference sequences
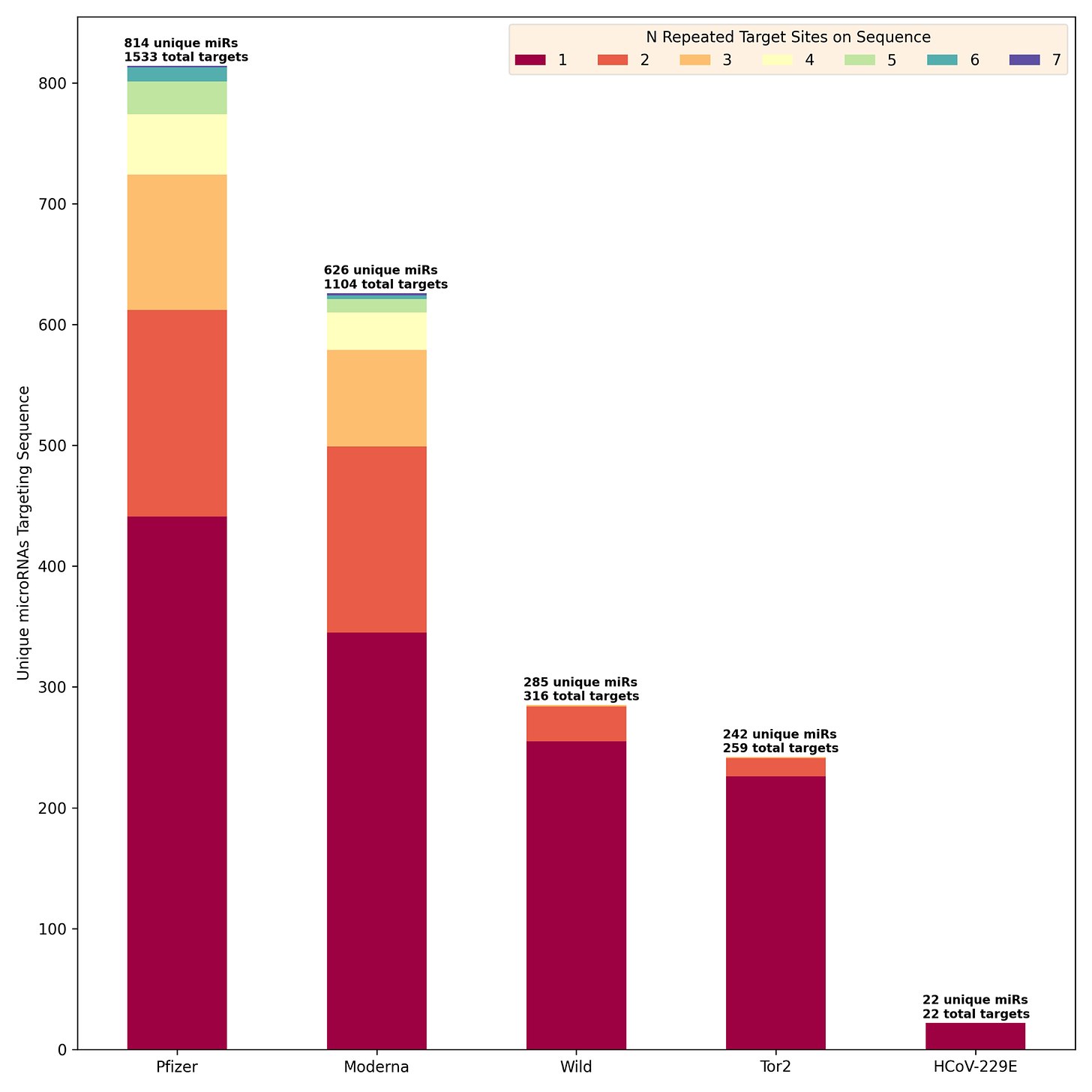
The figure below shows the number of unique microRNAs with one or more binding sites on each target sequence. The Pfizer and Moderna sequences are clearly targeted by many more microRNAs than any of the reference sequences. The Pfizer sequence targets 814 unique microRNAs having a total of 1533 binding sites and the Moderna sequence targets 626 unique microRNAs (1104 sites total), while the reference sequences are all under 300 unique microRNAs with about the same number of binding sites.
There is another interesting thing about this chart apart from the stark difference between Pfizer/Moderna microRNA counts and those of the reference sequences. Both Pfizer and Moderna sequences are targeted at multiple sites (up to 7) by a large number of microRNAs. This pattern is much less apparent in the SARS-CoV-2 wild strain and SARS-CoV Tor2, and it is not present at all in HCoV-229E. The chart below shows the counts of microRNAs targeting multiple sites more clearly. I’ve also listed the specific microRNAs having 5 or more repeated binding sites on the Pfizer (blue text) and Moderna (red text) below the chart. I find the progression from HCoV-229E - a presumably completely natural virus that has been around for a very long time - to SARS-CoV, SARS-CoV-2 and Pfizer/Moderna very interesting. The fact that HCoV-229E has a very low count of microRNA binding sites and no repeat sites suggests that there may be something “unnatural” about both the wild-type SARS-CoV-1/2 and the vaccine sequences.

If, in fact, the repeated binding sites are engineered, my speculation would be that these microRNAs are particularly important to sense (i.e. you want to make sure you don’t miss this signal so you add several sites on the spike mRNA to ensure that you catch it). However, it is also possible that the repeated sites could be useful for sponging host cell microRNAs to influence gene expression. Perhaps, both goals could be targeted for some of the microRNAs with repeated binding sites.
microRNA targeting by Pfizer/Moderna is statistically significant
The next question I asked myself was whether the difference between Pfizer/Moderna and the reference sequences was statistically significant. In order to answer this, I performed a Monte Carlo simulation where I analyzed 100 randomized Pfizer spike sequences for microRNA binding sites.

The results of this analysis showed that the number of microRNA binding sites on the Pfizer mRNA sequence is more than 5-sigma and the Moderna sequence more than 3-sigma from the distribution mean, while the wild-type Wuhan spike is around 0.5-sigma. We can conclude from this analysis that the Pfizer and Moderna sequences are statistically different from a randomized sequence in terms of microRNA binding sites.
Wild-type SARS-CoV-1/2 and Pfizer/Moderna preferentially target genes in the placenta and brain
After determining the microRNAs targeting the Pfizer/Moderna and reference sequences, the top 10 genes also targeted by each of those microRNAs was determined and then mRNA expression levels for those genes were obtained. The datasets were then grouped by gene and ranked in order of the total number of binding sites on the Pfizer/Moderna and reference sequences.
The heatmaps shown below show the top 50 genes ranked in order of number of binding sites on the vertical axis and tissue type on the horizontal axis. The plots to the right and top of each heatmap show the number of binding sites on the spike mRNA sequence targeting each gene and the total mRNA expression levels of all the genes by tissue type, respectively. The colors on the heatmap represent mRNA expression levels for particular combinations of genes and tissue types. Note also that some of the gene labels on the vertical axis are color-coded for the Pfizer and Moderna heatmaps. Green labels denote genes that are unique targets of both sequences, red labels denote genes that are unique to the Moderna sequence and blue labels denote those that are unique to Pfizer.

A general observation that is immediately evident from the heatmaps is that HCoV-229E is qualitatively different from all of the other sequences in that it appears to target microRNAs in a more random manner, which can be seen clearly in the expression plot above its heatmap. Conversely, the wild-type SARS-CoV-1/2 and Pfizer/Moderna spike sequences all target genes expressed in the placenta most heavily followed by brain tissues.
Differences among the SARS-CoV-1/2 and Pfizer/Moderna sequences appear to be relatively minor. One notable difference is the absence of targets for neutrophil genes in the Pfizer sequence. Also, the Pfizer sequence along with the SARS-CoV sequence appear to target the thalamus, whereas Moderna and SARS-CoV-2 do not particularly target it. Skeletal muscle tissues also appear to be heavily targeted by all of the sequences, except for perhaps SARS-CoV.
Another interesting feature can be seen in plots on the right side of the Pfizer and Moderna heatmaps. The top four genes targeted by these sequences are the same: NFIX, NFIC, IQSEC2 and SOGA1, although they have different rankings for the two sequences. Moreover, the number of target binding sites for these four genes appear to be relatively high compared to the remaining lower-ranking gene targets. In other words, there is an apparent step change in the number of binding sites upon moving from #4 to #5. This is particularly apparent for the Pfizer sequence.
The four genes NFIX, NFIC, IQSEC2 and SOGA1 are also conserved in the wild-type SARS-CoV-1/2 sequences. There are also five additional genes that are conserved across the wild-type and vaccine sequences: IGF2, ELK1, DAGLA, MECP2 and PPP19RB. The trends of the rankings of these conserved genes are curious, as you can see in the plot below. NFIC is consistently at the #1 or #2 spot for all sequences, while NFIX shows a steady progression from its lowest ranking in SARS-CoV (#17) to moderately higher rankings in SARS-CoV-2 (#5) and Moderna (#4) to the top spot in the Pfizer sequence. Remarkably, IQSEC2 and SOGA1 display similar trends. IGF2 moves in the opposite direction, starting at a ranking of #4 in the SARS-CoV sequence and moving up to #8 in Moderna and #19 in Pfizer. Perhaps, the most interesting gene in the chart is ELK1, which has a ranking of #35 in the SARS-CoV sequence, #2 in SARS-CoV-2, #3 in Moderna and back to #32 in Pfizer. DAGLA is also curious since its ranking in the SARS-CoV sequence (#23) and Moderna (#22) are roughly the same while also being the same for SARS-CoV-2 (#18) and Pfizer (#17). The other genes in the chart, MECP2 and PPP19RB seem to have a weak inverse correlation.
Finally, I want to point out the unique genes targeted by Pfizer and Moderna spike sequences. The Pfizer sequence uniquely targets FAM76A, AAK1, ZSWIM4, DNAJB5, TUBB4A and ZNF385A. The Moderna sequence targets LMLN, RERE, FBXO11, SDC3, SYT1, RAD51B, WDR48 and SCRT1. Both sequences uniquely target LIN28B, PRTG, HMGA2, IGDCC3, NR6A1, IGF2BP1, C14orf28, ARID3B, TRIM71, STARD13, PAX5, NFASC, IQSEC3 and MPP2. Generally speaking, you can see that the unique targets for each sequence are mostly confined to a ranking in the lower half of the top 50, while several of the shared unique genes appear clustered in the top 25 with the exceptions being PAX5, MPP2, IQSEC3 and NFASC which all occur in the lower 25 and in different orders for the two sequences.
Oh, and an important note - just because a gene is targeted by a sequence does not mean it will definitely result in its upregulation. First, we are relying on the results of calculations to identify binding sites. Although I have attempted to be as conservative as possible here, it is still possible that some microRNAs identified as having target sites may not in reality bind to their putative target sites. There is also the complicating factor of the Pfizer and Moderna mRNA vaccines using N1-methylpsuedouridine as a substitute for uridine in their sequences1. The RNA22v2 algorithm cannot account for this important difference, so it is possible that accuracy of the results might be impacted. However, as far as I can tell, the N1-methylpsuedouridine substitution results in stronger bonding to microRNAs, which would tend to suggest, if anything, that the RNA22v2 results might be too conservative.2 Another important issue to consider is that the microRNA, target sequence and host mRNA all have to be co-localized in the same cell in order for host gene expression to be altered. This fact should definitely be kept at the forefront of one’s mind when analyzing these results. Furthermore, it should be considered that the duration of any microRNA sponging effects mediated by Pfizer and Moderna sequences will be limited by the lifetime of their mRNA in the human body (currently unknown but most likely much longer than uridine-containing mRNA).
Discussion
In the discussion that follows my primary focus is to analyze the top gene targets of the Pfizer and Moderna spike sequences with a working assumption that these targets have been engineered into the sequences for the purpose of sponging microRNAs in host cells to influence gene expression.
I should remind the reader that there is another possible purpose for targeting a specific microRNA, which would be the detection of dangerous conditions in the host cell to shut-off spike protein translation. In fact, this may be the reason that repeated target sites (up to 7) for specific microRNAs have been added to the vaccine sequences. If the difference between life and death depends on detection of a single microRNA, then you want to make damn sure you are able to catch it. This topic surely deserves further attention.
At the end of this article, the reader will find a comprehensive review of most of the gene targets found in the following groups: 1) Conserved targets across SARS-CoV, SARS-CoV-2, Pfizer and Moderna; 2) Unique targets shared by Pfizer and Moderna; 3) Unique Pfizer targets; and 4) Unique Moderna targets.
While it is quite difficult to discern a clear pattern among all of these gene targets, there are some functional associations that appear to be meaningful:
Cardiac Repair - NFIC, IGF2, STARD13, DNAJB5
Cardiac Damage - NR6A1, ZSWIM4
Immune Suppression - NFIX, NFIC, ELK1, MECP2, LMLN
Immune Activation - DAGLA, PPP19RB, HMGA2, LIN28B, CLIP3, IGF2BP1, FBXO11, SDC3
Infection Facilitation - NFIC, IQSEC2/3, PRTG, IGF2BP1, AAK1
Adaptive Immunity - PAX5, ARID3B
Amyloid Formation - ELK1
Amyloid Suppression - DNAJB5
Mitigation of Spike-induced Effects - NFIX, NFIC, IGF2, ELK1, STARD13, PRTG, DNAJB5, RAD51B, WDR48
SARS-CoV-2 Interactors - HMGA2, ARID3B, IGF2BP1, TRIM71, SOGA1, NFIC, PPP1R9B, MECP2, WDR48, SYT1, TUBB4A, DNAJB5
It appears to me that the sequences examined here are a bit Janus-like in terms of the possible outcomes from microRNA-mediated upregulation of their target genes. Given the multiple roles that many proteins play in various circumstances, it should probably not be unexpected that both “good” and “evil” outcomes might be encountered. In the end, the effects of microRNA sponging will only become evident in the cells transfected by the virus/vaccine. Upregulation of a particular gene in a macrophage might have a very different outcome than the same gene being upregulated in, say, a neuron.
Perhaps the most surprising finding is that some of the genes potentially upregulated by microRNA sponging are those that have been shown to actively promote infection by SARS-CoV-2, in particular, AAK1 and IGF2BP1. Researchers have actually proposed therapeutics that suppress these genes to prevent infection. Is it possible that one goal of vaccination to seed the body with SARS-CoV-2 virions?
Finally, it is very curious to me that let-7-regulated genes are so heavily targeted by both the Pfizer and Moderna sequences. The suppression of these genes by let-7 is very important in adults since they are known to promote cancer. I’ve covered this in some detail in a previous article:
Conclusion
I must admit that there is a lot of information here and, honestly, it is beyond my ability to synthesize into a coherent framework. However, at a minimum, I feel that I have presented a compelling case that the Pfizer and Moderna vaccine sequences, and even the wild-type SARS-CoV and SARS-CoV-2 spike sequences, contain peculiar microRNA binding sites. In the case of Pfizer and Moderna, the sheer number of binding sites and the number of unique microRNAs with repeated binding sites is statistically significant. I’m convinced that there is something engineered here, even if I do not fully understand the purpose.
I encourage the reader to spend some time reviewing the information below to see if any patterns jump out that I may have missed. Also, I do have the spreadsheets containing the data presented in this article. I am happy to make them available to anyone who may have an interest. If there is sufficient interest from folks, I will consider posting them as an attachment.
The Conserved Gene Targets
NFIX
NFIX (Nuclear Factor 1 X-type) is a nuclear transcription factor that is expressed heavily in the brain, heart and skeletal muscle.
Involved with phenotype switching in macrophages
NFIX is also found in macrophages where it seems to play an important role in regulating the switch between pro- (M1) and anti-inflammatory (M2) phenotypes. One study found that macrophages lacking NFIX failed to adopt a M2 phenotype, while those expressing NFIX were able to do so3.
Significantly attenuates acute lung injury (ALI)
NFIX expression in lung epithelial tissues has been found to greatly reduce the damage from ALI caused by exposure to lipopolysaccharide (LPS). This outcome was attributed to NFIX-mediated down-regulation of NF-κB activation and decreased production of inflammatory factors such as TNF-α, IL-6 and IL-84.
Loss of circNFIX promotes cardiac repair following myocardial infarction
Cardiomyocyte proliferation in mice is inhibited by overexpression of circNFIX but promoted when knocked down5.
NFIC
NFIC (Nuclear Factor 1 C-type) is a nuclear transcription factor that is expressed heavily in the brain, heart and skeletal muscle.
Crucial importance in the response of the heart to ischemic cardiomyopathy
NFIC plays a very important role in the injured heart, downregulating TSPAN1, HOPX and possibly other genes. The genes regulated by NFIC mediate the activation of several damage-related pathways6.
Induces proliferation of myocytes
NFIC is significantly upregulated in acute myeloid leukemia. Overexpression of NFIC in hematopoietic stem and progenitor cells (HSPCs) was found to promote the differentiation of myocytes. Myocytes overexpressing NFIC were also found to have higher expression levels of growth and survival related genes7.
Identified as a crucial gene in SARS-CoV-2 infection of the GI tract
A bioinformatics study of gene regulatory network motifs identified NFIC as one of ten crucial genes involved with the infection of SARS-CoV-2 in the GI tract8.
Interacts with ORF14 protein of SARS-CoV-2 (see diagram above)
SOGA1
SOGA1 (Suppressor of Glucose, Autophagy-associated protein 1) is a protein that is widely expressed in the body, but predominately in the cerebellum. Very little is known about the possible functions of this protein.
Decreases glucose production and autophagy in response to insulin
SOGA1 participates in the insulin signaling pathway and mediates the reduction in glucose levels. SOGA1 also has an internal signaling peptide that can be released into circulation in response to insulin stimulation9.
Identified as a microtubule-associated protein
SOGA1 co-localizes with CLASP2 and tubulin forming a link between glucose metabolism and microtubules10.
Interacts with nsp7, nsp13 and nsp16 proteins of SARS-CoV-2 (see diagram above)
IQSEC2
IQSEC2 (IQ-motif and Sec7 domain 2) is a protein that is mostly expressed in the brain, but can also be found at high levels in skeletal muscle and the tongue. Mutations of this gene have been associated with X-linked intellectual disability.
Overexpression in neurons affects dendritic structure
Hippocampal cell cultures from mice that overexpressed IQSEC2 showed decreased axon length and simpler, more compact dendritic structures11.
Activates ARF6 which is an important protein for SARS-CoV-2 infection
IQSEC2 is a guanine nucleotide exchange factor which is known to activate ARF6 as its primary target12. ARF6 has been identified as a crucial membrane protein involved with the endocytic entry of SARS-CoV-2 into cells. Experiments using an ARF6 inhibitor showed a dose-dependent inhibition of viral infection13.
IGF2
IGF2 (Insulin-like growth factor 2) is a major growth factor expressed in the developing fetus, but also plays an important role in adults.
Upregulates VEGF expression in hypoxia
Kerationocytes exposed to IGF2 were found to upregulate expression of vascular endothelial growth factor (VEGF) and HIF-1α via the MAPK-ERK2 pathway14
Prevents retinal cell damage in the presence of reactive oxygen species
Retinal cells challenged with high levels of glucose were protected from apoptosis by overexpression of IGF215.
Promotes cardiac repair following myocardial infarction
Endothelial progenitor cells overexpressing IGF2 were used to treat rats with acute myocardial ischemia. IGF2 was found to promote proliferation of cardiomyocytes and suppress the inflammatory response and apoptosis16.
ELK1
ELK1 (ETS Like-1 protein) is an important transcription factor expressed ubiquitously in the body. It is the downstream target of the MAPK-ERK1/2 pathway and can form complexes with Serum Response Factor (SRF) to influence gene transcription.
Promotes ventilator-induced lung injury
ELK1 becomes upregulated during lung ventilation, leading to the downregulation of genes that are important for maintaining the integrity of cellular junctions17.
Promotes amyloid production in the brain
ELK1 has been found to be a potent repressor of presenilin-1 (PS1), a protein required for the proteolytic processing of amyloid precursor protein (APP). The inability to clear APP leads to accumulation of amyloid-β in the brain18. ELK1 has also been found to play an important role in Huntington’s disease, Parkinson’s disease and depression19.
Participates in IFN-I suppression in macrophages
The TPL-2 pathway is activated by TLR4 in macrophages and its downstream phosphorylation target is the ternary complex factor (TCF) formed by ELK1. The ELK1 TCF activated by this pathway upregulates FOS expression which leads to type I interferon suppression20.
May play a role in maintaining vascular integrity in response to SARS-CoV-2 spike protein
Myocardin (MYOCD) is known to compete with ELK1 for binding to SRF21. TGF-β stimulation of vascular smooth muscle cells can activate both MYOCD-SRF and ELK1-SRF via the MAPK/ERK1/2 pathway leading to a loss of differentiation in the former while the latter tends to upregulate growth and survival genes. The SARS-CoV-2 spike protein has been shown to induce barrier function and vascular leakage through interaction with the membrane integrins α5β1 which triggers TGF-β production22. Is it possible, then, that upregulation of ELK1 may help manage the cellular response to TGF-β induced by spike protein?
DAGLA
DAGLA (Diacylglycerol lipase alpha) is a protein expressed predominately in the brain. It is a serine hydrolase that is involved with the biosynthesis of endocannabinoids.
Promotes microglial activation in the brain
DAGLA-deficient mice challenged with LPS displayed reduced levels of neuroinflammation23.
MECP2
MECP2 (Methyl CpG binding protein 2) is a transcription factor that is widely expressed in the body, but is highly important in the nervous system where it is required for normal function of nerve cells.
Deficiency in brain has wide-ranging effects
MECP2 deficiency in microglia results in Rett-like symptoms while deficiency in astrocytes can spread to neurons and produce higher IL-1B and IL-6 levels in response to immune stimulation. MECP2 deficiency is also linked to microtubule instability in astrocytes and fibroblasts24.
Confers resilience to regulatory T-cells
MECP2 promotes resilience against inflammation to regulatory T-cells and protects against autoimmunity25
PPP1R9B
PPP1R9B (Neurabin-2, spinophilin) is a scaffold protein that is found throughout the body. It is highly expressed in the central nervous system.
Involved with immune synapse formation by NK-cells
PPP1R9B is recruited to the NK-cell immune synapse following engagement of CD18 and is critical for proper immune response26. CD18 is sharply downregulated by SARS-CoV-2 infection27.
Involved with endoplasmic reticulum stress response during SARS-CoV-2 infection28
Interacts with nsp1, nsp4 and nsp13 proteins of SARS-CoV-2 (see diagram above)
Gene Targets Shared by Pfizer and Moderna
PAX5
PAX5 (Paired box 5) is a transcription factor that plays an important role in B-cell maturation and in the development of the central nervous system and testis.
Promotes antibody class-switching
PAX5 stimulates the expression of the activation-induced cytidine deaminase (AID) gene in B-cells, which is essential for somatic hypermutation and antibody class-switching. PAX5 expression is required for optimal high-affinity IgG production29.
HMGA2
HMGA2 (High-mobility group AT-hook 2) is a transcription factor that is highly expressed in the developing fetus, but is strictly downregulated by let-7 in the adult body. Aberrant expression of HMGA2 in adults causes DNA damage and cancer.
Promotes inflammatory immune response
Human pulmonary alveolar epithelial cells challenged with LPS showed increased expression of HMGA2. Knockdown of HMGA2 resulted in reduced inflammation and apoptosis30.
Interacts with nsp2 protein of SARS-CoV-2 (see diagram above)
ARID3B
ARID3B (AT-rich interactive domain-containing protein 3B) is a DNA-binding protein that is highly expressed in the developing fetus, but is strictly downregulated by let-7 in the adult body.
Essential for B-cell development
Knockdown of ARID3B in the bone marrow of mice led to a reduction in B-cell population, while the populations of T-cells and other myeloid lineages were unchanged31.
Promotes apoptosis
ARID3B is a downstream transcriptional target of the p53 pathway that plays a key role in promoting the expression of pro-apoptotic genes32.
Interacts with nsp16 protein of SARS-CoV-2 (see diagram above)
LIN28B
LIN28B (Lin-28 homolog B) is a protein that is highly expressed in the testis, fetal liver and placenta. As an inhibitor of let-7 maturation, overexpression of this protein can result in the deregulation of let-7 suppressed genes, which can lead to cancer.
Promotes M1 macrophage phenotype
LIN28B knockdown in mice prevented Ang-II-mediated inhibition of let-7 formation which prevented M2 to M1 switch in macrophages33.
IGDCC3
IGDCC3 (Immunoglobulin Superfamily DCC Subclass Member 3) is a membrane protein that is not expressed widely in the adult body. It is thought to play a role in the neuromuscular process of maintaining balance. Very little is known about this protein.
Upregulated in covid-19 patients
IGDCC3 has been found to at elevated levels in nasal swabs taken from covid-19 patients34.
CLIP3
CLIP3 (CAP-Gly Domain Containing Linker Protein 3) is a cytoskeleton-associated protein that mediates the interaction of microtubules with organelles. It also plays a role in T-cell apoptosis.
Promotes astrocyte activation in response to injury
CLIP3 facilitates STAT3 signaling in spinal cord injury to activate astrocytes35.
STARD13
STARD13 (StAR-related lipid transfer domain protein 13) is a cytoskeleton-associated protein that is widely expressed throughout the body.
Protective against cardiotoxicity
Doxorubicin was found to cause cardiotoxicity by upregulating miR-125b which targets STARD13, suppressing the translocation of yes-associated protein from the nucleus to the cytoplasm36.
May rewire signaling pathways in SARS-CoV-2 infection
Study of covid-19 patients showed that STARD13 may have a slight protective effect37.
NR6A1
NR6A1 (nuclear receptor subfamily 6, group A, member 1), also known as Germ Cell Nuclear Factor, is a protein that is mainly expressed in the testis.
Promotes testicular tumor progression
NR6A1 is normally repressed by miR-196-5p but its downregulation leads to increased expression of NR6A1, which drives tumor growth38.
Associated with cardiac fibrosis in males
NR6A1 and NEK6 were both found to be highly expressed in the hearts of males with hypertrophic cardiomyopathy displaying fibrotic phenotype39.
PRTG
PRTG (Protogenin) is an immunoglobulin superfamily protein that is not widely expressed in the body.
Suppresses platelet aggregation
PRTG activates the cGMP/PKG signaling pathway which mediates the suppression of platelet aggregation40.
Activates membrane integrins that interact with spike protein
PRTG activates α5β1 integrins41. These integrins are known to interact with the spike protein of SARS-CoV-2 (see ELK1).
IGF2BP1
IGF2BP1 (Insulin-like growth factor 2 mRNA-binding protein 1) is a protein that binds to the mRNA of several important oncogenes, including IGF2. Its expression is strictly regulated in the by let-7 in the adult body.
Promotes M1 macrophage phenotype
Forced overexpression in macrophages and monocytes was found to promote an inflammatory response when challenged by LPS42.
Promotes SARS-CoV-2 infection
Knockdown of IGF2BP1 was found to reduce RNA levels of SARS-CoV-2 and ZIKV. IGF2BP1 directly binds to the SARS-CoV-2 RNA genome and promotes translation43.
Interacts with N, S, ORF3 and ORF9c proteins of SARS-CoV-2 (see diagram above)
IQSEC3 (see IQSEC2)
NFASC
NFASC (Neurofascin) is a protein that is highly expressed in the brain. It is a cell adhesion molecule involved with synapse formation in neurons.
Involved with endometriosis
NFASC has been found to be highly expressed in endometriosis and is correlated with increased infiltration of monocytes and M2 macrophages44.
May be involved with SARS-CoV-2 infection
NFASC was found to upregulated in the liver during SARS-CoV-2 infection45
TRIM71
TRIM71 (Tripartite Motif Containing 71) is an E3 ubiquitin-protein ligase that is expressed in the testis. It plays an important role in the developing fetus. It is also important for suppressing the expression of let-7.
Interacts with N protein of SARS-CoV-2 (see diagram above)
Gene Targets Unique to Pfizer
AAK1
AAK1 (Adaptor-associated protein kinase 1) is a protein that is expressed widely in the adult body. It plays an important role in regulating clathrin-mediated endocytosis.
Promotes SARS-CoV-2 infection
Endocytosis is one way in which SARS-CoV-2 virions can enter the cell. Knockdown of AAK1 has been found to inhibit SARS-CoV-2 entry by endocytosis. Therapeutics have been proposed which target AAK1 inhibition to prevent SARS-CoV-2 infection46.
ZSWIM4
ZSWIM4 (Zinc Finger SWIM-Type Containing 4) is a protein that is not widely expressed in the adult body. It is thought to enable zinc ion binding and to be a part of the CuI2-RING complex.
May promote ischemic injury of the heart
As part of the CuI2-RING ubiquitin ligase complex, ZSWIM4 promotes the ubiquitination and degradation of SMAD147. SMAD1 mediates cellular signaling triggered by TGF-β which is produced following myocardial injury. SMAD1 has been suggested to play a protective role in ischemia through effects that may involve the activation of anti-apoptotic signals48. It is possible then, that ZSWIM4 may worsen cardiac damage by degradation of SMAD-1.
DNAJB5
DNAJB5 (DnaJ Heat Shock Protein Family (Hsp40) Member B5) is a heat-shock protein that is expressed heavily in the heart.
Promotes cardiac regeneration after injury
DNAJB5 is a critical protein involved with cardiac regeneration after injury49. It also acts as a master negative regulator of cardiac hypertrophy.
Suppresses amyloid formation
DNAJB5 and other members of this family have been found to be potent inhibitors of cytotoxic protein aggregation50.
Interacts with M protein of SARS-CoV-2 (see diagram above)
TUBB4A
TUBB4A (Tubulin beta-4A chain) is tubulin protein that is expressed heavily in the brain. It forms microtubules in the cytoplasm by bonding with Tubulin-α.
Involved with Acute Respiratory Distress Syndrome (ARDS) caused by SARS-CoV-2
ARDS patients were found to have high levels of TUBB4A along with low levels of neutrophils51. SARS-CoV-2 is thought to “hijack” the microtubule cytoskeleton during infection through interactions between its spike protein and the microtuble wall52.
Interacts with nsp10, nsp13 and ORF3 proteins of SARS-CoV-2 (see diagram above)
ZNF385A
ZNF385A (Zinc Finger Protein 385A) is a zinc finger protein that is expressed heavily in the brain.
Promotes cell cycle arrest in response to DNA damage
ZNF385A is induced by p53 and binds to its DNA binding domain which leads to preferential activation of pro-arrest genes rather than pro-apoptotic genes53. Cationic lipid nanoparticles have been shown to cause significant DNA damage54.
Gene Targets Unique to Moderna
LMLN
LMLN (Leishmanolysin-like (metallopeptidase M8 family)), also known as invadolysin, is a metalloprotease that is not widely expressed in the adult body. It is known to play an important role in the virulence of the parasite Leishmania.
Interacts with lipid particles
LMLN is associated with the surface of lipid particles. The role that it plays in this context is unclear, but it is speculated to influence the dynamics of lipid particles and perhaps have a functional role as well55.
Promotes immune evasion by parasites
Parasites such as Schistosoma mansoni deploy invadolysin to suppress the innate immune system. Treatment of leukocytes with invadolysin has been shown to reduce the release of key immunostimulatory cytokines56.
FBXO11
FBXO11 (F-box only protein 11) is a protein that is expressed throughout the body. It is one of four proteins in a ubiquitin ligase complex which acts to degrade target proteins in a phosphorylation-dependent manner.
Promotes IFN-I signaling in response to viral RNA/DNA
FBXO11 amplifies IFN-I signaling by promoting the phosphorylation of TBK1 and IRF357.
SDC3
SDC3 (Syndecan-3) is a membrane protein that is expressed throughout the body. It is involved in the organization and assembly of the extracellular matrix.
Promotes inflammatory response in endothelial cells
SDC3 is highly expressed in endothelial cells where it plays a role in chemokine presentation at the cellular membrane and coordination of leuokocyte trafficking58.
SYT1
SYT1 (Synaptotagmin-1) is a membrane expressed predominately in the brain where it is involved with vesicular trafficking and exocytosis.
Promotes cell-to-cell spread of viruses
SYT1 regulates endocytosis of viruses and enhances cell-to-cell spread via the endocytic recycling pathway59.
Interacts with ORF3a protein of SARS-CoV-2 protein (see diagram above)
RAD51B
RAD51B (RAD51 Paralog B) is a DNA repair protein found throughout the body.
WDR48
WDR48 (WD Repeat Domain 48) is a protein that promotes deubiquitinating activity.
Promotes DNA repair
WDR48 binds with RAD51B to promote the repair of damaged DNA60.
Interacts with S, M and ORF3a proteins of SARS-CoV-2 (see diagram above)
Kim KQ, Burgute BD, Tzeng SC, Jing C, Jungers C, Zhang J, Yan LL, Vierstra RD, Djuranovic S, Evans BS, Zaher HS. N1-methylpseudouridine found within COVID-19 mRNA vaccines produces faithful protein products. Cell Rep. 2022 Aug 30;40(9):111300.
Parr CJC, Wada S, Kotake K, Kameda S, Matsuura S, Sakashita S, Park S, Sugiyama H, Kuang Y, Saito H. N 1-Methylpseudouridine substitution enhances the performance of synthetic mRNA switches in cells. Nucleic Acids Res. 2020 Apr 6;48(6):e35.
Saclier M, Lapi M, Bonfanti C, Rossi G, Antonini S, Messina G. The Transcription Factor Nfix Requires RhoA-ROCK1 Dependent Phagocytosis to Mediate Macrophage Skewing during Skeletal Muscle Regeneration. Cells. 2020 Mar 13;9(3):708.
Zhang XL, An J, Deng YZ, Fang XZ, Xu CY, Liu XF, Bai ZH, Zhang G, Cui MY. A novel miRNA-762/NFIX pathway modulates LPS-induced acute lung injury. Int Immunopharmacol. 2021 Nov;100:108066.
Huang S, Li X, Zheng H, Si X, Li B, Wei G, Li C, Chen Y, Chen Y, Liao W, Liao Y, Bin J. Loss of Super-Enhancer-Regulated circRNA Nfix Induces Cardiac Regeneration After Myocardial Infarction in Adult Mice. Circulation. 2019 Jun 18;139(25):2857-2876.
Ye Y, Jin Q, Gong Q, Li A, Sun M, Jiang S, Jin Y, Zhang Z, He J, Zhuang L. Bioinformatics and Experimental Analyses Reveal NFIC as an Upstream Transcriptional Regulator for Ischemic Cardiomyopathy. Genes (Basel). 2022 Jun 13;13(6):1051.
Rastogi N, Gonzalez JBM, Srivastava VK, Alanazi B, Alanazi RN, Hughes OM, O'Neill NS, Gilkes AF, Ashley N, Deshpande S, Andrews R, Mead A, Rodrigues NP, Knapper S, Darley RL, Tonks A. Nuclear factor I-C overexpression promotes monocytic development and cell survival in acute myeloid leukemia. Leukemia. 2023 Feb;37(2):276-287.
Ghani S, Kalantari S, Mirmotalebisohi SA, Sameni M, Poursheykhi H, Dadashkhan S, Abbasi M, Zali H. Specific Regulatory Motifs Network in SARS-CoV-2-Infected Caco-2 Cell Line, as a Model of Gastrointestinal Infections. Cell Reprogram. 2022 Feb;24(1):26-37.
Cowherd RB, Asmar MM, Alderman JM, Alderman EA, Garland AL, Busby WH, Bodnar WM, Rusyn I, Medoff BD, Tisch R, Mayer-Davis E, Swenberg JA, Zeisel SH, Combs TP. Adiponectin lowers glucose production by increasing SOGA. Am J Pathol. 2010 Oct;177(4):1936-45.
Kruse R, Krantz J, Barker N, Coletta RL, Rafikov R, Luo M, Højlund K, Mandarino LJ, Langlais PR. Characterization of the CLASP2 Protein Interaction Network Identifies SOGA1 as a Microtubule-Associated Protein. Mol Cell Proteomics. 2017 Oct;16(10):1718-1735.
Hinze SJ, Jackson MR, Lie S, Jolly L, Field M, Barry SC, Harvey RJ, Shoubridge C. Incorrect dosage of IQSEC2, a known intellectual disability and epilepsy gene, disrupts dendritic spine morphogenesis. Transl Psychiatry. 2017 May 2;7(5):e1110.
Levy NS, Umanah GKE, Rogers EJ, Jada R, Lache O, Levy AP. IQSEC2-Associated Intellectual Disability and Autism. Int J Mol Sci. 2019 Jun 21;20(12):3038.
Mirabelli C, Sherman EJ, Cunha JB, Wotring JW, El Saghir J, Harder J, Kretzler M, Sexton JZ, Emmer BT, Wobus CE. ARF6 is a host factor for SARS-CoV-2 infection in vitro. bioRxiv [Preprint]. 2022 Nov 7:2022.
Kwon YW, Kwon KS, Moon HE, Park JA, Choi KS, Kim YS, Jang HS, Oh CK, Lee YM, Kwon YG, Lee YS, Kim KW. Insulin-like growth factor-II regulates the expression of vascular endothelial growth factor by the human keratinocyte cell line HaCaT. J Invest Dermatol. 2004 Jul;123(1):152-8.
Zhao Y, Xiong Z, Chen Y, Wang G, Zhao Y. Activation of Insulin-Like Growth Factor-2 Ameliorates Retinal Cell Damage and Exerts Protection in in vitro Model of Diabetic Retinopathy. Neuroimmunomodulation. 2021;28(4):266-275.
Demetz G, Oostendorp RAJ, Boxberg AM, Sitz W, Farrell E, Steppich B, Steinsiek AL, Rudelius M, Ott I. Overexpression of Insulin-Like Growth Factor-2 in Expanded Endothelial Progenitor Cells Improves Left Ventricular Function in Experimental Myocardial Infarction. J Vasc Res. 2017;54(6):321-328.
Tao Z, Jie Y, Mingru Z, Changping G, Fan Y, Haifeng W, Yuelan W. The Elk1/MMP-9 axis regulates E-cadherin and occludin in ventilator-induced lung injury. Respir Res. 2021 Aug 23;22(1):233.
Pastorcic M, Das HK. Ets transcription factors ER81 and Elk1 regulate the transcription of the human presenilin 1 gene promoter. Brain Res Mol Brain Res. 2003 May 12;113(1-2):57-66.
Besnard A, Galan-Rodriguez B, Vanhoutte P, Caboche J. Elk-1 a transcription factor with multiple facets in the brain. Front Neurosci. 2011 Mar 16;5:35.
Blair L, Pattison MJ, Chakravarty P, Papoutsopoulou S, Bakiri L, Wagner EF, Smale S, Ley SC. TPL-2 Inhibits IFN-β Expression via an ERK1/2-TCF-FOS Axis in TLR4-Stimulated Macrophages. J Immunol. 2022 Feb 15;208(4):941-954.
Wang, Z., Wang, DZ., Hockemeyer, D. et al. Myocardin and ternary complex factors compete for SRF to control smooth muscle gene expression. Nature 428, 185–189 (2004).
Biering SB, de Sousa FTG, Tjang LV, Pahmeier F, Ruan R, Blanc SF, Patel TS, Worthington CM, Glasner DR, Castillo-Rojas B, Servellita V, Lo NTN, Wong MP, Warnes CM, Sandoval DR, Clausen TM, Santos YA, Ortega V, Aguilar HC, Esko JD, Chui CY, Pak JE, Beatty PR, Harris E. SARS-CoV-2 Spike triggers barrier dysfunction and vascular leak via integrins and TGF-β signaling. bioRxiv [Preprint]. 2021 Dec 13:2021.12.10.472112. doi: 10.1101/2021.12.10.472112. Update in: Nat Commun. 2022 Dec 9;13(1):7630.
Viader A, Ogasawara D, Joslyn CM, Sanchez-Alavez M, Mori S, Nguyen W, Conti B, Cravatt BF. A chemical proteomic atlas of brain serine hydrolases identifies cell type-specific pathways regulating neuroinflammation. Elife. 2016 Jan 18;5:e12345.
Jin XR, Chen XS, Xiao L. MeCP2 Deficiency in Neuroglia: New Progress in the Pathogenesis of Rett Syndrome. Front Mol Neurosci. 2017 Oct 4;10:316.
Li C, Jiang S, Liu SQ, Lykken E, Zhao LT, Sevilla J, Zhu B, Li QJ. MeCP2 enforces Foxp3 expression to promote regulatory T cells' resilience to inflammation. Proc Natl Acad Sci U S A. 2014 Jul 8;111(27):E2807-16.
Meng X, Kanwar N, Du Q, Goping IS, Bleackley RC, Wilkins JA. PPP1R9B (Neurabin 2): involvement and dynamics in the NK immunological synapse. Eur J Immunol. 2009 Feb;39(2):552-60.
Barros-Martins J, Förster R, Bošnjak B. NK cell dysfunction in severe COVID-19: TGF-β-induced downregulation of integrin beta-2 restricts NK cell cytotoxicity. Signal Transduct Target Ther. 2022 Jan 31;7(1):32.
https://pathway-viewer.toolforge.org/embed/WP4861
Nasri Nasrabadi P, Martin D, Gharib E, Robichaud GA. The Pleiotropy of PAX5 Gene Products and Function. Int J Mol Sci. 2022 Sep 3;23(17):10095.
Shi L, Shi CW, Cheng KW. HMGA2 Synergizes with EZH2 to Mediate Epithelial Cell Inflammation and Apoptosis in Septic Lung Dysfunction. Ann Clin Lab Sci. 2022 Nov;52(6):938-946.
Kurkewich JL, Klopfenstein N, Hallas WM, Wood C, Sattler RA, Das C, Tucker H, Dahl R, Cowden Dahl KD. Arid3b Is Critical for B Lymphocyte Development. PLoS One. 2016 Aug 18;11(8):e0161468.
Pratama E, Tian X, Lestari W, Iseki S, Ichwan SJ, Ikeda MA. Critical role of ARID3B in the expression of pro-apoptotic p53-target genes and apoptosis. Biochem Biophys Res Commun. 2015 Dec 4-11;468(1-2):248-54.
Jaiswal A, Maurya M, Maurya P, Barthwal MK. Lin28B Regulates Angiotensin II-Mediated Let-7c/miR-99a MicroRNA Formation Consequently Affecting Macrophage Polarization and Allergic Inflammation. Inflammation. 2020 Oct;43(5):1846-1861.
https://www.frontiersin.org/articles/10.3389/fcimb.2022.1011672/full
Chen X, Chen C, Hao J, Zhang J, Zhang F. Effect of CLIP3 Upregulation on Astrocyte Proliferation and Subsequent Glial Scar Formation in the Rat Spinal Cord via STAT3 Pathway After Injury. J Mol Neurosci. 2018 Jan;64(1):117-128.
Jin X, Yu W, Ye P. MiR-125b enhances doxorubicin-induced cardiotoxicity by suppressing the nucleus-cytoplasmic translocation of YAP via targeting STARD13. Environ Toxicol. 2022 Apr;37(4):730-740.
Pellegrina D, Bahcheli AT, Krassowski M, Reimand J. Human phospho-signaling networks of SARS-CoV-2 infection are rewired by population genetic variants. Mol Syst Biol. 2022 May;18(5):e10823.
Liu X, Fan Z, Li Y, Li Z, Zhou Z, Yu X, Wan J, Min Z, Yang L, Li D. microRNA-196a-5p inhibits testicular germ cell tumor progression via NR6A1/E-cadherin axis. Cancer Med. 2020 Dec;9(23):9107-9122.
Xu F, Chen Y, Tillman KA, Cui Y, Williams RW, Bhattacharya SK, Lu L, Sun Y. Characterizing modifier genes of cardiac fibrosis phenotype in hypertrophic cardiomyopathy. Int J Cardiol. 2021 May 1;330:135-141.
Zhang F, Mu G, Liu Z, Xie Q, Zhang H, Zhou S, Wang Z, Hu K, Wang Z, Zhao X, Cui Y, Xiang Q. Genetic Polymorphisms Associated with Prothrombin Time and Activated Partial Thromboplastin Time in Chinese Healthy Population. Genes (Basel). 2022 Oct 15;13(10):1867.
Wang YC, Juan HC, Wong YH, Kuo WC, Lu YL, Lin SF, Lu CJ, Fann MJ. Protogenin prevents premature apoptosis of rostral cephalic neural crest cells by activating the α5β1-integrin. Cell Death Dis. 2013 Jun 6;4(6):e651.
Xie J, Li Q, Zhu XH, Gao Y, Zhao WH. IGF2BP1 promotes LPS-induced NFκB activation and pro-inflammatory cytokines production in human macrophages and monocytes. Biochem Biophys Res Commun. 2019 Jun 11;513(4):820-826.
Zhang, S., Huang, W., Ren, L. et al. Comparison of viral RNA–host protein interactomes across pathogenic RNA viruses informs rapid antiviral drug discovery for SARS-CoV-2. Cell Res 32, 9–23 (2022).
Chen P, Yao M, Fang T, Ye C, Du Y, Jin Y, Wu R. Identification of NFASC and CHL1 as Two Novel Hub Genes in Endometriosis Using Integrated Bioinformatic Analysis and Experimental Verification. Pharmgenomics Pers Med. 2022 Apr 22;15:377-392.
Hammoudeh SM, Hammoudeh AM, Bhamidimarri PM, Al Safar H, Mahboub B, Künstner A, Busch H, Halwani R, Hamid Q, Rahmani M, Hamoudi R. Systems Immunology Analysis Reveals the Contribution of Pulmonary and Extrapulmonary Tissues to the Immunopathogenesis of Severe COVID-19 Patients. Front Immunol. 2021 Jun 28;12:595150.
Ghamry, H.I., Belal, A., El-Ashrey, M.K. et al. Evaluating the ability of some natural phenolic acids to target the main protease and AAK1 in SARS COV-2. Sci Rep 13, 7357 (2023).
Chengdong Wang, Ziran Liu, Yelin Zeng et al. ZSWIM4 regulates early embryonic patterning and BMP signaling pathway by promoting nuclear Smad1 degradation, 13 July 2022, PREPRINT (Version 1)
Hanna A, Humeres C, Frangogiannis NG. The role of Smad signaling cascades in cardiac fibrosis. Cell Signal. 2021 Jan;77:109826. doi: 10.1016/j.cellsig.2020.109826. Epub 2020 Nov 5.
Pang M, Xiong C, Xiao C, Du J, Zheng L, Bai L, Zhu X, Xiong JW. Critical role of zebrafish dnajb5 in myocardial proliferation and regeneration. J Genet Genomics. 2020 Aug;47(8):493-496.
Hageman J, Rujano MA, van Waarde MA, Kakkar V, Dirks RP, Govorukhina N, Oosterveld-Hut HM, Lubsen NH, Kampinga HH. A DNAJB chaperone subfamily with HDAC-dependent activities suppresses toxic protein aggregation. Mol Cell. 2010 Feb 12;37(3):355-69.
García-Hidalgo MC, Peláez R, González J, Santisteve S, Benítez ID, Molinero M, Perez-Pons M, Belmonte T, Torres G, Moncusí-Moix A, Gort-Paniello C, Aguilà M, Seck F, Carmona P, Caballero J, Barberà C, Ceccato A, Fernández-Barat L, Ferrer R, Garcia-Gasulla D, Lorente-Balanza JÁ, Menéndez R, Motos A, Peñuelas O, Riera J, Bermejo-Martin JF, Torres A, Barbé F, de Gonzalo-Calvo D, Larráyoz IM. Genome-wide transcriptional profiling of pulmonary functional sequelae in ARDS- secondary to SARS-CoV-2 infection. Biomed Pharmacother. 2022 Oct;154:113617. doi: 10.1016/j.biopha.2022.113617. Epub 2022 Aug 30.
Aminpour M, Hameroff S, Tuszynski JA. How COVID-19 Hijacks the Cytoskeleton: Therapeutic Implications. Life (Basel). 2022 May 30;12(6):814.
Das S, Raj L, Zhao B, Kimura Y, Bernstein A, Aaronson SA, Lee SW. Hzf Determines cell survival upon genotoxic stress by modulating p53 transactivation. Cell. 2007 Aug 24;130(4):624-37. doi: 10.1016/j.cell.2007.06.013.
Zhanataev AK, Anisina EA, Kulakova AV, Shilovskiy IP, Lisitsyn AA, Koloskova OO, Khaitov MR, Durnev AD. Genotoxicity of cationic lipopeptide nanoparticles. Toxicol Lett. 2020 Aug 1;328:1-6.
Cobbe N, Marshall KM, Gururaja Rao S, Chang CW, Di Cara F, Duca E, Vass S, Kassan A, Heck MM. The conserved metalloprotease invadolysin localizes to the surface of lipid droplets. J Cell Sci. 2009 Sep 15;122(Pt 18):3414-23. doi: 10.1242/jcs.044610. Epub 2009 Aug 25.
Hambrook JR, Hanington PC. A cercarial invadolysin interferes with the host immune response and facilitates infection establishment of Schistosoma mansoni. PLoS Pathog. 2023 Feb 2;19(2):e1010884.
Gao L, Gao Y, Han K, Wang Z, Meng F, Liu J, Zhao X, Shao Y, Shen J, Sun W, Liu Y, Xu H, Du X, Li J, Qin FX. FBXO11 amplifies type I interferon signaling to exert antiviral effects by facilitating the assemble of TRAF3-TBK1-IRF3 complex. J Med Virol. 2023 Mar;95(3):e28655.
Eustace AD, McNaughton EF, King S, Kehoe O, Kungl A, Mattey D, Nobbs AH, Williams N, Middleton J. Soluble syndecan-3 binds chemokines, reduces leukocyte migration in vitro and ameliorates disease severity in models of rheumatoid arthritis. Arthritis Res Ther. 2019 Jul 12;21(1):172.
Lewis JD, Lazarowitz SG. Arabidopsis synaptotagmin SYTA regulates endocytosis and virus movement protein cell-to-cell transport. Proc Natl Acad Sci U S A. 2010 Feb 9;107(6):2491-6.
https://www.genecards.org/cgi-bin/carddisp.pl?gene=WDR48&keywords=wdr48










This is very interesting stuff. Are these differences the result of different codon optimization, or are they in untranslated regions?
If it is in the optimization, it could be that the virus in nature evolves to avoid these binding sites, leaving apparent optimizations available to engineer.
Or it could be more sinister.